EBRAINS at Imagine ArtScience 2024

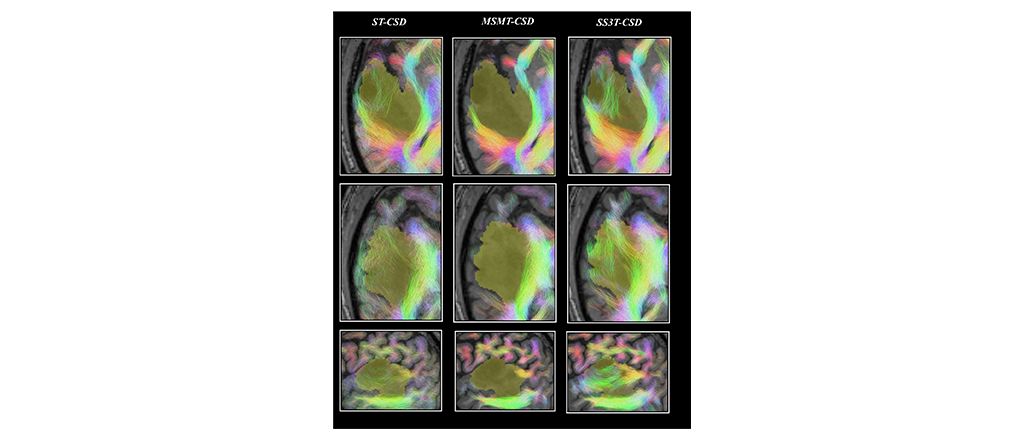
Comparison of tractography results based on single tissue (ST)-CSD, Multi Shell Multi Tissue (MSMT)-CSD and Single Shell 3 Tissues (SS3T-CSD)34 pipelines for patient PAT26 (anaplastic astrocytoma WHO grade III)
In 2020, it was estimated that over 300,000 people were diagnosed with a primary brain or spinal cord tumor worldwide. Brain tumors can be treated with either surgery, radiation therapy, or chemotherapy. In tumor resection surgery, a surgeon aims to remove as much of a tumor as possible, while preserving brain tissue and overall function.
Although surgery is often able to reduce tumor mass, halt progression, and preserve neurological function, outcomes are difficult to predict. To alleviate some of this uncertainty, there is growing interest in using personalized brain models of patients, which would allow clinicians to preview the effects of surgery.
An international team of researchers has taken the first step in a proof of concept of modeling brains with lesions in the first place, and then performing virtual surgeries on them by collecting functional MRI data from brain tumor patients at the University of Ghent. Data was collected from 11 glioma patients, 14 meningioma patients, and 11 healthy caregivers. Glioma tumors originate from glial cells, while meningioma tumors (which account for around thirty percent of brain tumor cases), are found in the membranes that surround the brain and spinal cord. The neuroimaging data was collected in two phases - first, the day before each patients’ surgery, and then, approximately six months later.
The data has since been used for brain simulations of tumor patients before and after surgery in The Virtual Brain - an open-source platform for constructing and simulating personalised brain network models on EBRAINS. The full fMRI dataset can also be accessed on the EBRAINS Knowledge Graph. Crucially, the data contains important derivatives, such as the estimated shape of the hemodynamic response function (the mapping between neural activity and signal recorded by fMRI) for each brain region and subject, as well as the parameters of the computational models, already stored according to the Brain Imaging Data Standard, making them seamlessly interoperable and reusable.
“I am particularly happy about this dataset, which reflects different stages of my research path, and different aspects of neuroinformatics and neuroscience research in Europe,” said Daniele Marinazzo, one of the lead researchers. “It all started with the PhD project of Hannelore Aerts, in which she used The Virtual Brain to build individual models of brain tumor patients using fMRI data. A few years later, our group joined the Human Brain Project and contributed to the integration of the Virtual Brain in the EU neuroscience flagship. Due to this conjecture, the timing with respect to changing GDPR rules, and the excellent work of the data curators, this dataset was among the first on the platform. Thanks to the efforts of Prof. Ritter’s group at Charité Hospital, we were able to include the parameters of the computational models in BIDS standards. Part of this effort were also a couple of excellent Google Summer of Code students, recruited through the International Neuroinformatics Coordination Facility.” He added: “Last but not least, in order to acknowledge the crucial role of hemodynamics in personalized models of BOLD signal, in particular in brain tumors, we adapted the TVB framework to include a proxy of the resting state hemodynamic response function. I am confident that other users will join us in this effort (some of the additions to the dataset came already from the feedback of some colleagues interested in working on the data). The recent integration of macroscale and microscale models, now possible in EBRAINS, will definitely allow a deeper exploration of the pathophysiology of brain tumors from a computational perspective.”


EBRAINS is open and free. Sign up now for complete access to our tools and services.