3DSpineMFE
A MATLAB® toolbox that given a three-dimensional spine reconstruction computes a set of characteristic morphological measures that unequivocally determine the spine shape.
BioExcel Building Blocks Workflows is a collection of biomolecular workflows to explore the flexibility and dynamics of macromolecules, including signal transduction proteins or molecules related to the Central Nervous System. Molecular dynamics setup for protein and protein-ligand complexes are examples of workflows available as Jupyter Notebooks. The workflows are built using the BioBB software library, developed in the framework of the BioExcel Centre of Excellence. BioBBis a collection of Python wrappers on top of popular biomolecular simulation tools, offering a layer of interoperability between the wrapped tools, which make them compatible and prepared to be directly interconnected to build complex biomolecular workflows.
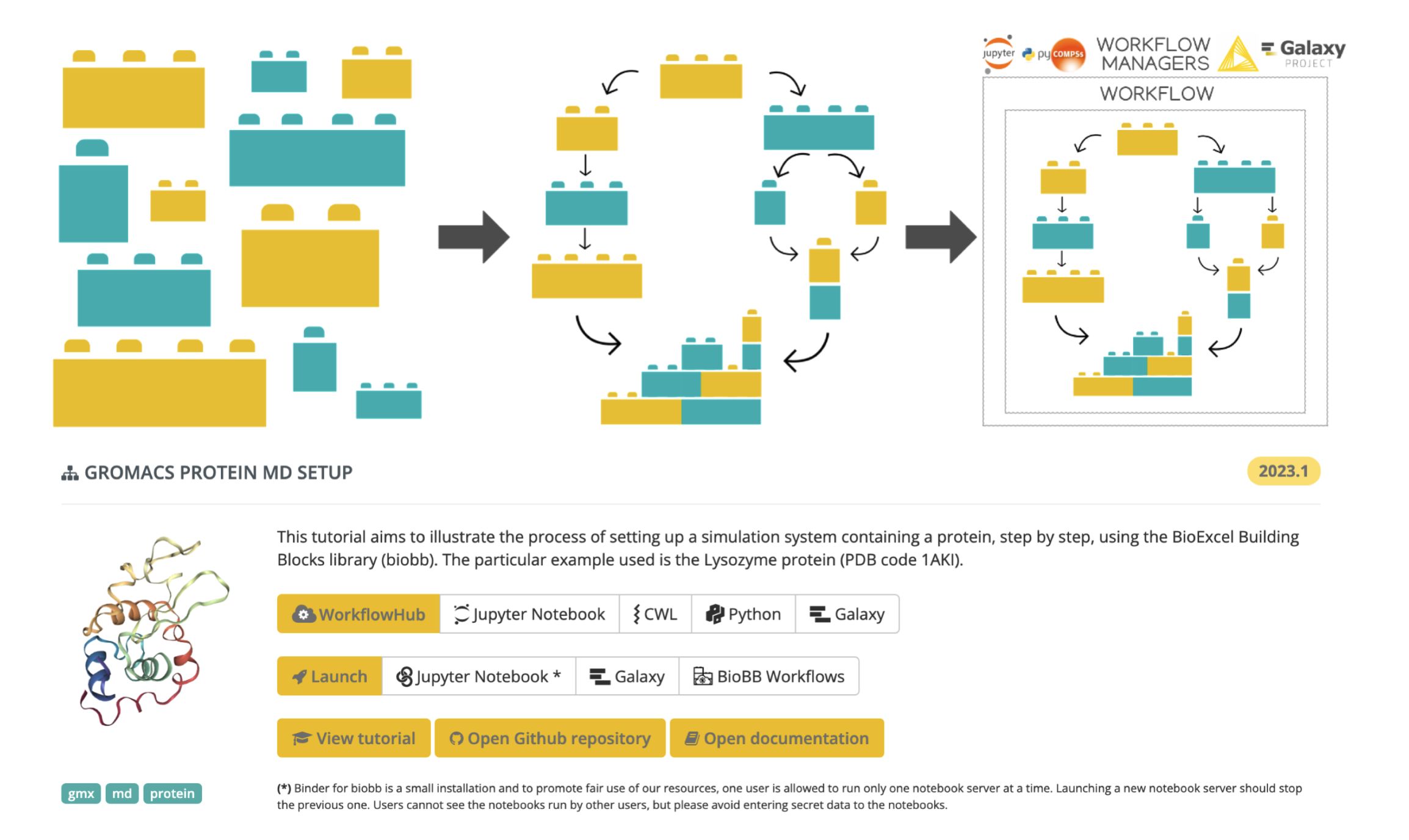
BioExcel Building Blocks (BioBB) library giving interoperability to biomolecular simulation tools, and associated workflows (e.g. Protein MD setup with GROMACS MD engine).
Publication:
Andrio, P., Hospital, A., Conejero, J. et al. BioExcel Building Blocks, a software library for interoperable biomolecular simulation workflows. Sci Data 6, 169 (2019). https://doi.org/10.1038/s41597-019-0177-4
A MATLAB® toolbox that given a three-dimensional spine reconstruction computes a set of characteristic morphological measures that unequivocally determine the spine shape.
Arbor is a high-performance library for computational neuroscience simulations with multi-compartment, morphologically-detailed cells, from single cell models to very large networks. Arbor is written from the ground up with many-cpu and gpu architectures in mind, to help neuroscientists effectively use contemporary and future HPC systems to meet their simulation needs. Arbor supports NVIDIA and AMD GPUs as well as explicit vectorization on CPUs from Intel (AVX, AVX2 and AVX512) and ARM (Neon and SVE). When coupled with low memory overheads, this makes Arbor an order of magnitude faster than the most widely-used comparable simulation software. Arbor is open source and openly developed, and we use development practices such as unit testing, continuous integration, and validation.
ArDock employs the arbitrary docking method to reveal potential interaction sites on the surface of a protein by computationally docking a set of random protein “probes”. The random probes interact in a non-random manner on protein surfaces, and the targeted regions are enriched in biological interfaces. Docking is performed on input protein structures using the Hex software. The ArDock webserver performs the docking calculations and provides tools for the combined analysis of protein structures and sequences and for the visualization of the results to identify interaction sites.
EBRAINS is open and free. Sign up now for complete access to our tools and services.