The Virtual Brain
Create personalised brain models and simulate multi-scale networks.
- Ready-to-go image-processing pipelines to generate structural and functional data for personalised brain models.
- Simulation-ready brain image data.
- TVB-NEST co-simulation software to simulate the brain at multiple scales.
- Extra-fast optimised TVB code for high-performance computing.
The Virtual Brain (TVB) is an open-source platform for constructing and simulating personalised brain network models. The TVB-on-EBRAINS ecosystem includes a variety of prepackaged modules, integrated simulation tools, pipelines and data sets for easy and immediate use on EBRAINS. Process your large cohort databases and use these results to develop potential medical treatments, therapies or diagnostic procedures.
Access TVB in the Cloud or download and run it elsewhere
Run TVB simulations remotely from your local computer via the TVB EBRAINS Cloud simulator with HPC back end (free EBRAINS account required) or download the TVB Docker container or TVB standalone (OS-compiled), via the TVB website.
See the TVB Collaboratory for tips on how EBRAINS can help you maximise the TVB resources.
Launch TVB EBRAINS Cloud Simulator Download TVB Standalone from the TVB website Download TVB Container Get access to the TVB Collaboratory
Simulate the brain at multiple scales with TVB-Multiscale Co-Simulation
Couple large-scale brain models from TVB with spiking networks from NEST Simulator (link to tool page for NEST Simulator). For example, you may study the impact of realistic large-scale brain network activity on spiking neurons and networks, or even replace entire brain areas by spiking networks and study emerging mean field dynamics.
Access the multi-scale features of TVB in one click via the Web App. Easily set up and run TVB-NEST workflows, choosing Python wrapper integration or TVB-NEST-Elephant co-simulation. TVB-Multiscale Co-Simulation is also available for download as a Docker container.
Launch TVB-Multiscale Simulation Web App
Download TVB-multiscale Container
Related Resources
See all resourcesTVB on EBRAINS: The Virtual Brain integrated workflows on EBRAINS (Long Version)
TVB on EBRAINS: short videos
The Virtual Brain: INCF TVB Training Space
TVB-NEST integration wrapper
TVB-NEST integration wrapper via the Collaboratory
Watch TVB-NEST-Elephant workflow
TVB-NEST-Elephant co-simulation workflow
Increase simulation speed using FAST-TVB
Exploit parallel supercomputing architectures with multi-threaded TVB simulations to perform faster-than-real-time simulations of large networks with FAST-TVB. You can use the convenient platform-independent Docker container on your computer or supercomputer.
How to use the Fast-TVB
TVB workflow for inferring a spatial map of the hyperexcitable zone in patient brains with epilepsy
Use TVB in a sophisticated model inversion approach: uncover the model’s secret parameter values and identify the zone with hyperexcitable parameters; the epileptogenic zone, by applying the statistical methods used in the EPINOV trial.
This approach combines best-in-class simulator technology with the best available inference techniques and is extensible, enabling its application on other pathologies. A set of Jupyter notebooks takes you step-by-step through the entire workflow.
TVB-inverse
Process your MRI data sets with the TVB Image Processing Pipeline
The TVB Image Processing Pipeline takes multi-modal MRI data sets (anatomical, functional or diffusion MRI) as input. Your output includes structural and functional connectomes, region-average fMRI time series, brain surface triangulations, EEG electrode positions, lead field matrices and brain parcellations. The pipeline takes data sets in BIDS standard format and gives outputs that can be used for further analysis or to construct personalised brain models. In addition, outputs comply with the BIDS standards, are annotated with metadata following the MINDS schema and are adapted for sharing via the EBRAINS Knowledge Graph. Run our Jupyter notebook example to see how the TVB processing pipeline can be executed from the web browser.
TVB - EBRAINS integrated workflows: Image Processing Pipeline
Find TVB ready-processed imaging data in the EBRAINS Knowledge Graph
Use processed TVB ready brain scans to run your simulations. This reference data set contains a series of brain scans from a total of 25 tumour patients and 11 healthy controls, organised in the BIDS format.
Find the data set in the Knowledge Graph
Interactive Atlas Viewer in Multiple Languages
A Virtual Standard Brain model and its network parcellations are available in an interactive atlas viewer. Explore brain parcellations used by TVB, the functional relevance of different brain areas and networks and their location and shapes. Select areas based on either anatomy or function. Descriptions are available in English, Arabic, German, Hebrew and Polish.
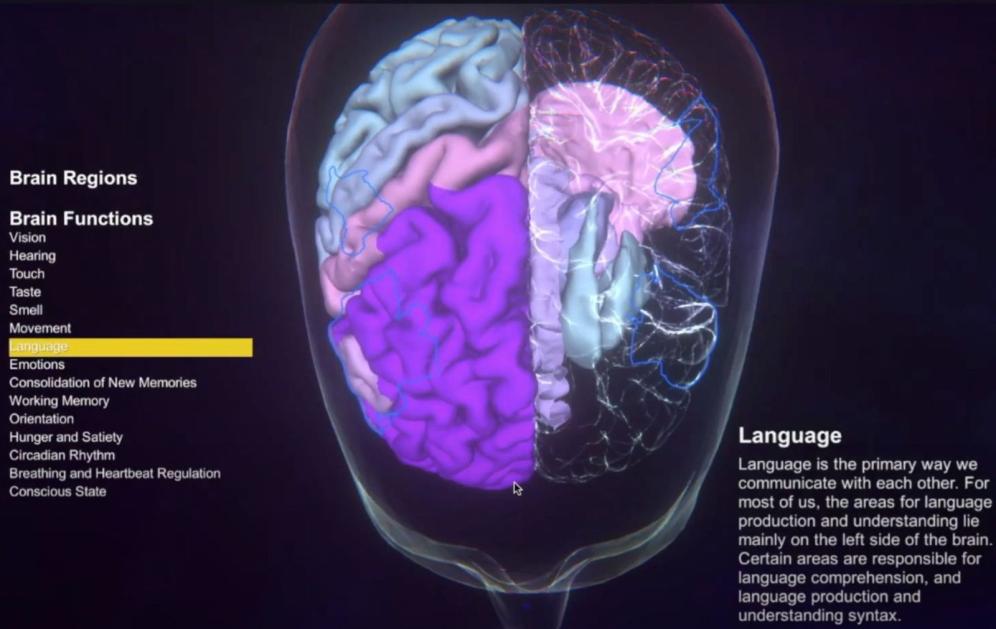
Brain Atlas Viewer
How to use TVB-on-EBRAINS
The Virtual Brain integration in EBRAINS allows users to have an end-to-end experience of personalised brain model creation and multi-scale brain simulation, using high-performance computing in the Cloud. This makes it possible to process large-cohort databases and produce generalised results - a precondition for using these results to develop potential medical treatments, therapies or diagnostic procedures.
How to run:
a) Run pre-installed software in the Cloud with web access and HPC back end.
b) Pull execution-ready Docker containers from Docker Hub.
c) Download software or execution-ready containers and run them on a system of your choice, including your local machine.
- Execute the containers on EBRAINS HPC, using secure container environments like Sarus or Shifter.
- Operate the containers on supercomputers directly from a web browser via Jupyter notebooks.
- Log in directly to a shell of the supercomputer via SSH and run the software.
d) Download software or execution-ready containers and run them on a system of your choice, including your local machine.
Short video about TVB on EBRAINS
Full video about TVB on EBRAINS
Mini videos explaining TVB on EBRAINS
INCF TVB Training Space
Other Tools
See all toolsCreate an account
EBRAINS is open and free. Sign up now for complete access to our tools and services.